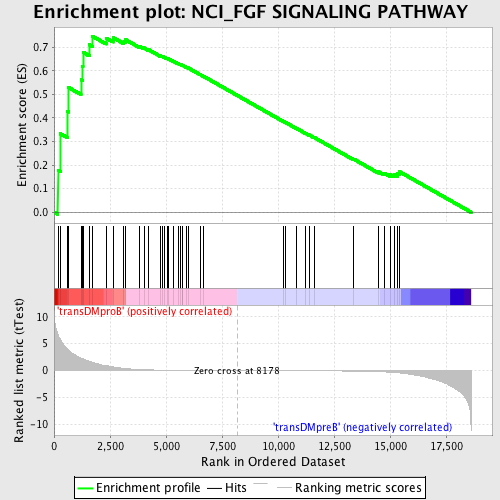
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_transDMpreB.phenotype_transDMproB_versus_transDMpreB.cls #transDMproB_versus_transDMpreB.phenotype_transDMproB_versus_transDMpreB.cls #transDMproB_versus_transDMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_transDMpreB.cls#transDMproB_versus_transDMpreB_repos |
| Upregulated in class | transDMproB |
| GeneSet | NCI_FGF SIGNALING PATHWAY |
| Enrichment Score (ES) | 0.7468141 |
| Normalized Enrichment Score (NES) | 1.6279391 |
| Nominal p-value | 0.0018450185 |
| FDR q-value | 0.17378901 |
| FWER p-Value | 0.378 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | STAT1 | 3936 5524 | 173 | 6.741 | 0.1774 | Yes | ||
| 2 | GAB1 | 18828 | 280 | 5.825 | 0.3331 | Yes | ||
| 3 | PDPK1 | 23097 | 588 | 4.023 | 0.4280 | Yes | ||
| 4 | MAPK1 | 1642 11167 | 646 | 3.755 | 0.5289 | Yes | ||
| 5 | JUN | 15832 | 1203 | 2.302 | 0.5628 | Yes | ||
| 6 | PMF1 | 12452 | 1266 | 2.180 | 0.6198 | Yes | ||
| 7 | PAK4 | 17909 | 1291 | 2.157 | 0.6783 | Yes | ||
| 8 | SPP1 | 5501 | 1578 | 1.710 | 0.7103 | Yes | ||
| 9 | PTK2B | 21776 | 1704 | 1.562 | 0.7468 | Yes | ||
| 10 | MMP9 | 14732 | 2326 | 0.900 | 0.7383 | No | ||
| 11 | GRB2 | 20149 | 2633 | 0.675 | 0.7405 | No | ||
| 12 | FGFR2 | 1917 4722 1119 | 3116 | 0.398 | 0.7256 | No | ||
| 13 | FOS | 21202 | 3200 | 0.360 | 0.7311 | No | ||
| 14 | PLAU | 22084 | 3818 | 0.187 | 0.7031 | No | ||
| 15 | FGF1 | 1994 23447 | 4015 | 0.150 | 0.6967 | No | ||
| 16 | SSH1 | 10540 | 4193 | 0.124 | 0.6906 | No | ||
| 17 | CDH1 | 18479 | 4746 | 0.076 | 0.6630 | No | ||
| 18 | MAPK3 | 6458 11170 | 4819 | 0.071 | 0.6611 | No | ||
| 19 | SHC1 | 9813 9812 5430 | 4928 | 0.066 | 0.6571 | No | ||
| 20 | HGF | 16916 | 5055 | 0.061 | 0.6520 | No | ||
| 21 | PIK3R1 | 3170 | 5108 | 0.059 | 0.6508 | No | ||
| 22 | MET | 17520 | 5322 | 0.052 | 0.6408 | No | ||
| 23 | FGFR4 | 3226 | 5533 | 0.046 | 0.6308 | No | ||
| 24 | SOS1 | 5476 | 5642 | 0.043 | 0.6261 | No | ||
| 25 | PIK3CA | 9562 | 5729 | 0.040 | 0.6226 | No | ||
| 26 | SPRY2 | 21725 | 5747 | 0.040 | 0.6228 | No | ||
| 27 | IL17RD | 22069 | 5925 | 0.036 | 0.6143 | No | ||
| 28 | STAT5B | 20222 | 5983 | 0.035 | 0.6121 | No | ||
| 29 | PTPN11 | 5326 16391 9660 | 6513 | 0.024 | 0.5843 | No | ||
| 30 | NCAM1 | 5149 | 6688 | 0.022 | 0.5756 | No | ||
| 31 | SRC | 5507 | 10226 | -0.026 | 0.3859 | No | ||
| 32 | CAMK2A | 2024 23541 1980 | 10344 | -0.028 | 0.3803 | No | ||
| 33 | PLCG1 | 14753 | 10828 | -0.036 | 0.3553 | No | ||
| 34 | SDC2 | 9134 | 11217 | -0.043 | 0.3356 | No | ||
| 35 | FGFR1 | 3789 8968 | 11386 | -0.046 | 0.3279 | No | ||
| 36 | RUNX2 | 4480 8700 | 11615 | -0.052 | 0.3170 | No | ||
| 37 | AKT1 | 8568 | 13376 | -0.114 | 0.2254 | No | ||
| 38 | CTTN | 8817 4575 1035 8818 | 14473 | -0.221 | 0.1725 | No | ||
| 39 | FGF23 | 17275 | 14728 | -0.259 | 0.1660 | No | ||
| 40 | CBL | 19154 | 15001 | -0.315 | 0.1601 | No | ||
| 41 | CDH2 | 1963 8727 8726 4508 | 15181 | -0.372 | 0.1608 | No | ||
| 42 | RPS6KA1 | 15725 | 15336 | -0.422 | 0.1642 | No | ||
| 43 | PLAUR | 18351 | 15414 | -0.446 | 0.1724 | No |

